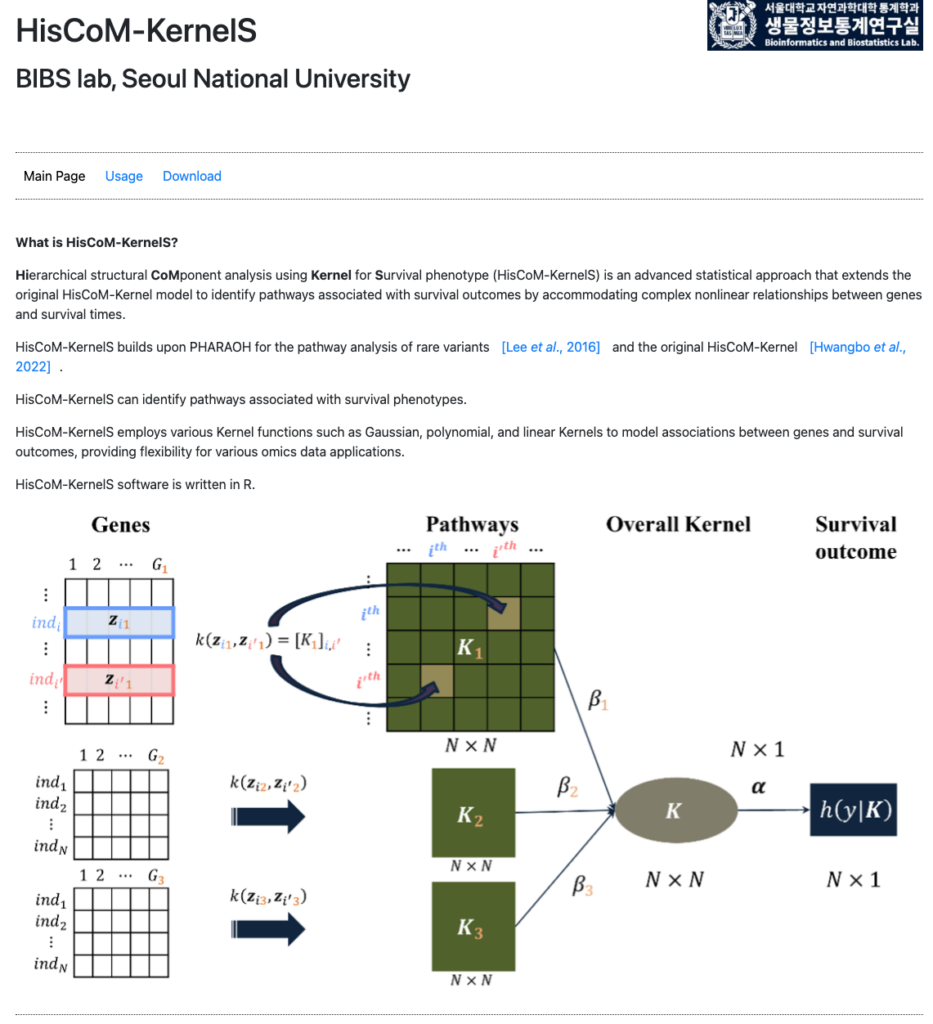

link: http://statgen.snu.ac.kr/software/HisCom-KernelS/

link: http://statgen.snu.ac.kr/software/HisCom-KernelS/
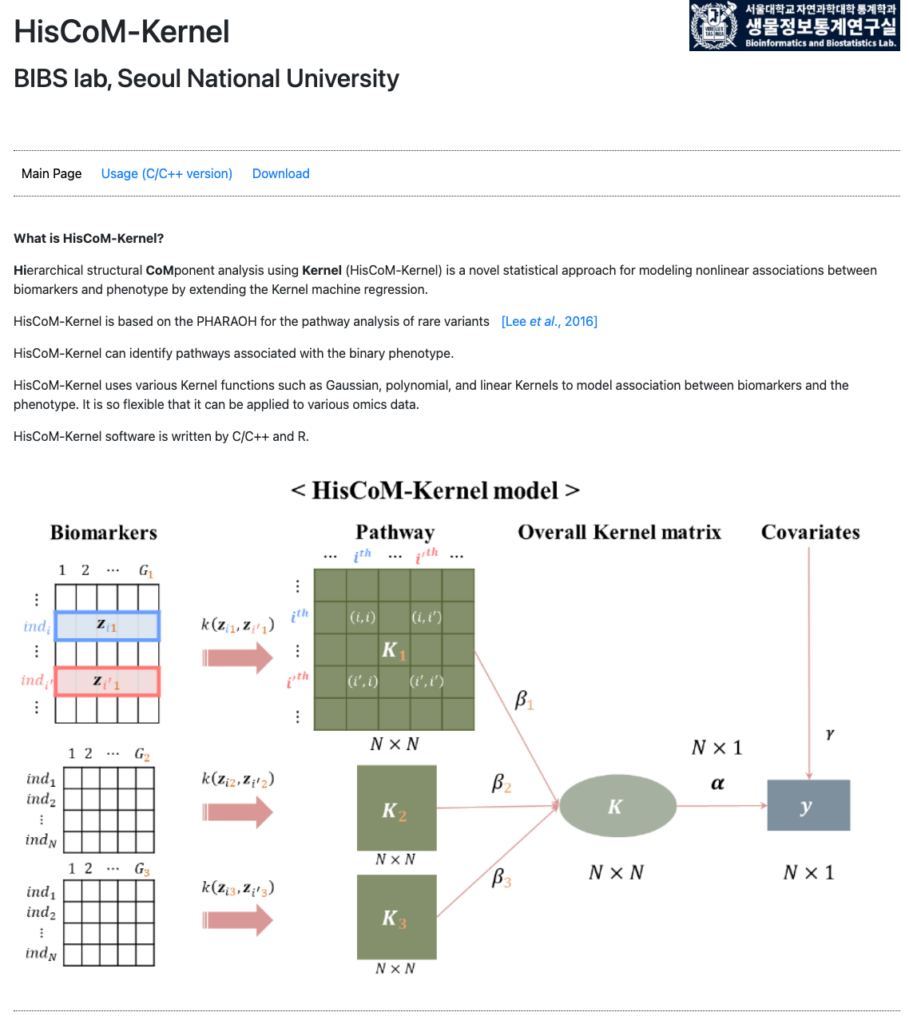

link: http://statgen.snu.ac.kr/software/HisCom-Kernel/
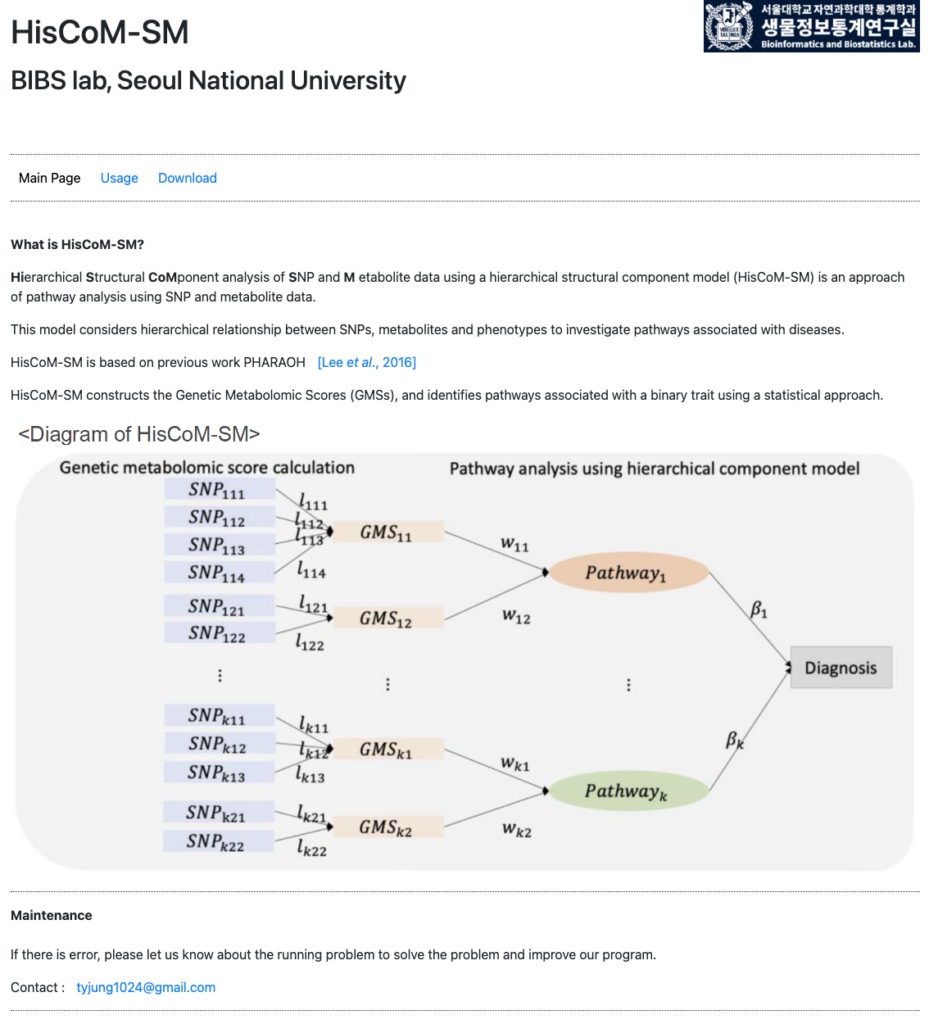

link: http://statgen.snu.ac.kr/software/HisCoM_SM/

link: http://statgen.snu.ac.kr/software/HisCom-PAGE/
link: http://statgen.snu.ac.kr/software/HisCoM-Categ.html

link: http://statgen.snu.ac.kr/software/HisCom-PCA/

link: http://statgen.snu.ac.kr/software/hiscom-mimi/